Welcome to the RNA/ssDNA pulling server !
This server performs quantitative
predictions of force-extension
curves of RNA or ssDNA
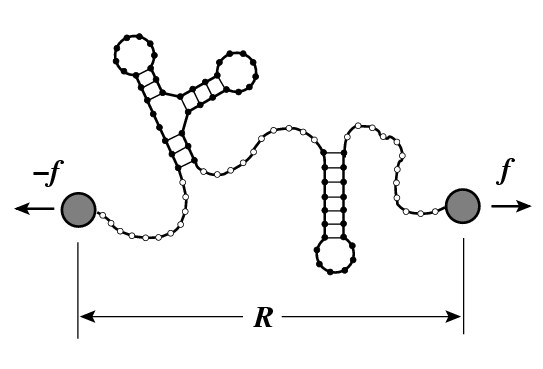
molecules. It takes the secondary
structure of the molecule fully
into account with the exception
of pseudoknots. The single-stranded
pieces of the molecule are modeled
as an elastic freely jointed chain.
|
|
Back to the Bundschuh group's
server page or to the
mirror at
Terry Hwa's group.
If you want to know more about how the software behind this server works,
please read the article
Ulrich Gerland, Ralf Bundschuh, and Terence Hwa,
Force-induced denaturation of RNA,
Biophys. J. 81 (2001),
1324-1332.
The secondary-structure prediction part of this software is an extension
of the Vienna
package described in
I.L. Hofacker, W. Fontana, P.F. Stadler, S.Bonhoeffer, M. Tacker,
and P. Schuster, Fast folding and comparison of RNA secondary
structures, Monatshefte f. Chemie 125 (1994)
167-188.
For questions and/or comments please send an email to rnapuller@bioserv.mps.ohio-state.edu.
Back to the Bundschuh group's
server page.
11/23/18, Ralf
Bundschuh, The Ohio State
University
